In the last post, we saw how we can use Pipelines to streamline our machine learning workflow.
We will start off with a fantastic feature that will blow the lid off your mind. All the effort done till now, in post I and II, can be done in 2 lines of code! Ah yes, this magic is possible with use of a feature called AutoML. Not only will it perform preprocessing steps automatically, it will also select the best algorithm from a multitude of them including XGBoost, LightGBM, Prophet etc. The hyper parameter search comes for free 🙂 If all this was not enough it will share the entire auto-generated code for your use and modification. So, the two magic lines are:
from databricks import automl
summary = automl.classify(train, target_col="category", timeout_minutes=20)
The AutoML requires you to specify the following:
- Type of machine learning problem – Classification/ Regression/ Forecasting
- Specify the training set, target column
- Timeout after which the autoML will stop looking for better models.
While all this is very exciting, the real world use case of AutoML tends to be to create a baseline for model performance and give us a model to start with. In our case, the AutoML gave an RoC score of 91.8% (actually better than our manual work so far!!) for the best model.
Looking at the auto-generated Python notebook, here are the broad steps it took:
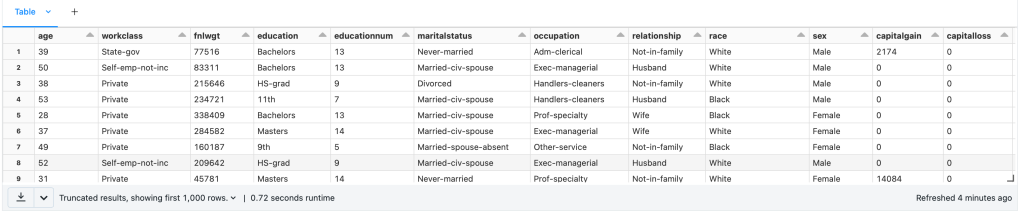
- Load Data
- Preprocessing
- Impute values for missing numerical columns
- Convert each categorical column into multiple binary columns through one-hot encoding
- Train – Validation – Test Split
- Train classification model
- Inference
- Determine Accuracy – Confusion matrix, ROC and Precision-Recall curves for validation data
This is broadly in line with what we did in our manual setup.
Apart from the coded approach we saw, you can create AutoML experiment using the UI from the Experiments -> Create AutoML Experiment button. If you can’t find the experiments tab, make sure that you have the Machine Learning persona selected in the Databricks workspace.
In an enterprise/ real world scenario, we will build many alternate models with different parameters and use the one with higest accuracy. Next, we deploy the model and use it for predictions with new data. Finally, the selected model will be updated over time as new data becomes available. Until now, we didn’t talk about how to handle these enterprise requirements or were doing it manually on best effort.
These requirements are covered under what we know as MLOps. Spark supports MLOps using an open source framework called MLFlow. MLFlow supports the following objects:
- Projects: Provides a mechanism for storing machine learning code in a reusable and reproducible format.
- Tracking: Allows for tracking your experiments and metrics. You can see the history of your model and its accuracy evolve over time. There is also a tracking UI available.
- Model Registry: Allows for storing various models that you develop in a registry with an UI to explore the same. It also provides for model lineage (which MLflow experiment and run produced the model), stage transitions (for example from staging to production)
- Model Serving: We can use serving to provide inference endpoints for either batch or inline processing. Mostly this will be made available as REST endopoints.
There is a very easy way to get started with MLFlow where we allow MLFlow to log automatically the metrics and models. This can be done using a single line:
import mlflow
mlflow.autolog()This will log the parameters, metrics, models and the environment. The core concept to get with MLOps is the concept of runs. Each run is a unique combination of parameters and algorithm that you have executed. Many runs can be part of the same experiment.
To get started we can set name of the experiment with the command: mlflow_set_experiment(“name_of_the_experiment”)
To start tracking the experiments manually, we can setup the context as follows:
with mlflow.start_run(run_name="") as run:
<pseudo code for running an experiment>
mlflow.log_param("key","value")
mlflow.log_metrics("key","value")
mlflow.spark.log_model(model,"model name")
You can track parameters and metrics using log_param and log_metrics functions of mlflow object. The model can now be registered with the Model Registry using the function: mlflow.register_model(model_uri=””, name=””)
What is important here is the model_uri. The model_uri takes the form: runs:/<runid>/model. The runid identifies the specific run in the experiment and each model is stored at the model_uri location mentioned above.
You can now load the model and perform inference using the following code:
import mlflow
# Load model
loaded_model = mlflow.spark.load_model(model_uri)
# Perform inference via model.transform()
loaded_model.transform(data)
While we have seen how to track experiments explicitly, Databricks Workspaces also track the experiments automatically (from Databricks Runtime 10.3 ML and above). You can view the expriments and their runs in the UI via the Experiments sidebar of the Machine Learning persona.
You will need to click on the specific experiment (in our case Adult Dataset), that will show all the runs of the experiment. Click on the specific run to get more details about the run. The run will show the metrics recorded which is in our case was areaUnderRoC of 91.4%.
Under Artifacts, if you click on the model, you can see the URI of the model run. This URI can be used to register the model with the Model Registry and use it for predictions at any point in time.
MLFlow also supports indicating the state of the model for production. Different states supported by MLFlow are:
- None
- Staging
- Production
- Archived
Once your model is registered with the Model registry, you can change the state of the model to any other state with the function transition_model_version_stage() function.
From the model registry you are able to create model serving endpoint using the Serverless Real-Time Inference service that uses managed Databricks compute service to provide a REST endpoint.